At the core of every drug development program is the challenge to successfully express active and high-quality proteins. Drug developers often are faced with protein-expression issues, struggling to express a specific protein for months (and sometimes years) with varying degrees of success. Low or no expression, insoluble expression, and proteolytic clipping of an expressed protein or a combination thereof are examples of the hurdles encountered in product development efforts.
With protein expression being vital to the success of every stage of biopharmaceutical development, there is an inherent need to screen as many variables as rapidly as feasible to find optimal production strain and fermentation conditions. Focused on the rapid development of biologics, Pfenex Inc. has spent years specializing in high-throughput and parallel screening for strain engineering and process development. Using Pfēnex expression technology, a Pseudomonas fluorescens– based platform, Pfenex can rapidly screen thousands of expression strains to identify the optimal production strain (1). The platform has been constructed to assess the performance of thousands of host strain/plasmid combinations in parallel to identify a production strain for a specific protein of interest. Host-strain phenotypes that are available off the shelf include those with one or more deletions in protease genes, strains capable of coexpressing folding modulator proteins, and disulfide bond isomerases, along with combinations of these.
PRODUCT FOCUS: ANTIBODIES, VACCINES, BIOSIMILARS
PROCESS FOCUS: PRODUCTION
WHO SHOULD READ: PRODUCT DESIGN, PROCESS ENGINEERS
KEYWORDS: MICROBIAL EXPRESSION, HIGH-THROUGHPUT SCREEENING, LABEL-FREE ASSAY
LEVEL: INTERMEDIATE
With advantages in quality, stability, solubility, and titer, Pfenex technology platform reduces production and development costs facilitating the rapid delivery of products through preclinical and clinical development and into commercialization (2). This platform technology enables Pfenex's own pipeline of biosimilars, novel biotherapeutics, and vaccines. In addition to fueling its own product pipeline, the company also leverages its technology platform through Strain Engineering and Reagent Proteins divisions to support others in the research community, as well as those developing their own therapeutics and vaccines.
Pfenex's technology platform rapidly accelerates product development by assuring first-time, significant production of high-quality, active, and soluble protein within five weeks. Specific components of the platform were designed to ensure soluble and active expression through avoidance of proteolytic clipping and posttranslational modifications along with enhancements to solubility through protein-folding improvements. Genetic control elements on expression plasmids (promoters, ribosome binding sites, secretion leaders) have been combined to generate >150 off-the-shelf, unique, rapid-cloning vectors, allowing control of high-quality protein expression (3).
Predicting which components will be crucial in successful expression of a given protein cannot be determined a priori from intrinsic information such as the amino acid sequence. The combination of critical factors is empirical for each individual target protein. Accordingly, Pfenex has developed a high-throughput, robotically enabled screening work process in 96-well format. The process is coupled to high-throughput analytical methods, which allow the parallel evaluation of thousands of unique expression strains containing a variety of expression strategies for a specific gene.
Such observation and quantitation of specific protein expression is essential for identification of expression strains for further development and optimization of bioreactor titers in production. Traditional analytical methods that have been used for this analysis include sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), enzyme-linked immunosorbent assay (ELISA), and high-performance liquid chromatography (HPLC). Those methods have certain drawbacks, including poor accuracy and precision as well as laborious methods such as data analysis. Conversely, label-free assays offer a solution for bioprocessing bottlenecks, while addressing product quality.
One such label-free assay is bio-layer interferometry (BLI), used by the family of Octet instruments from ForteBio (Pall). Those instruments support Pfenex's high-throughput and rapid parallel screening of expression strains. Their analytical capabilities have advantages where existing methods such as HPLC and ELISA have limitations in sample throughput and performance. With full-plate protein quantitation in 15–30 minutes of 96 samples, Octet systems provide direct label-free quantitation of antibodies or other proteins that are important for early expression strain screening and biologics manufacturing. Biosensors can measure protein concentrations in the presence of extraneous matter such as cell debris, which is typically present in cell lysate and fermentation cell-free broth, thus providing rapid titer measurements without labor-intensive sample preparation and purification.
BLI technology monitors the binding of proteins and other biomolecules to their binding partners/receptors directly, in real time. Binding interaction is continuously monitored by measuring the change in thickness of the protein layer on the tip of a biosensor (4). The method does not require labeling of the protein with fluorescent or chromogenic tags, thus eliminating interference issues.
Dozens of protein-specific BLI methods have been developed at Pfenex for a number of protein types expressed in P. fluorescens. Off-the-shelf biosensor chemistries include protein A, antihuman and antimurine IgG, antipenta-HIS, and streptavidin. Pfenex often generates its own custom biosensors protein by protein. That typically involves identifying and obtaining a high-purity binding partner (e.g., protein) for the target protein being expressed. The binding partner is then biotinylated, typically through NHS-ester crosslinking, and then bound to streptavidin biosensors immediately before use.
Case Study
Using its core technology, Pfenex developed a high-producing production strain for expression of soluble granulocyte colony stimulating factor (G-CSF) (5). The currently marketed nonglycosylated form of G-CSF is produced in Escherichia coli as an inclusion body that requires solubilization and refolding (6). Pfenex's G-CSF is expressed properly folded and soluble for low-cost production as a biosimilar in the therapeutic protein market.
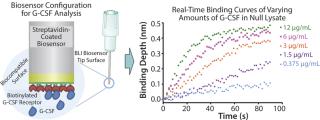
The first stage in developing a production process for G-CSF was to use Pfenex's expression platform and tool box to screen hundreds of expression strains to identify those strains that were capable of producing soluble, correctly folded G-CSF. The Octet system with a G-CSF receptor immobilized to streptavidin biosensors was used for titer measurements. This method demonstrated sufficient range and sensitivity for detecting soluble, functional G-CSF directly with minimal sample preparation and low interference from null lysate (Figure 1).
Figure 1:
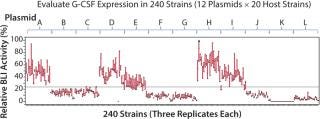
Pfenex used BLI to screen in parallel 240 unique expression strains. Twelve plasmids carrying the G-CSF gene fused to different P. fluorescens secretion leaders were constructed, targeting the protein to the periplasm. The plasmids were transformed into 20 phenotypically unique host strains and the soluble fractions of replicate culture lysates were analyzed for G-CSF receptor binding using the BLI method. The resulting screen identified several dozen expression strains that exhibited high soluble expression levels of G-CSF (Figure 2).
Figure 2:
As commonly observed using the Pfenex platform, secretion leader selection had a significant influence on the level of expression in each expression strain. Three secretion leaders showed high expression. Of the remaining secretion leaders, six showed less relative G-CSF expression, two showed medium expression (Figure 2), and one secretion leader did not enable soluble G-CSF expression. Approximately 30 expression strains were selected on the basis of specific expression levels and diversity of host and secretion leaders for further evaluation (data not shown). The entire process to identify a production strain took less than five weeks. The BLI assay provided both a means to assess the amount of soluble G-CSF produced by the expression strains as well as an indication of the functional quality of the material by its ability to bind to the G-CSF receptor in the initial screening process. Subsequently, the five highest-producing expression strains were selected for fermentation range finding.
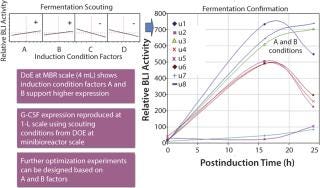
First, five expression strains were tested in a design of experiments (DoE) format testing different fermentation conditions at a 4-mL scale. Pfenex also was able to use the same G-CSF BLI assay during this fermentation-range–finding stage. BLI analysis revealed fermentation conditions that positively affected G-CSF expression level (Figure 3).
Figure 3:
Subsequently, we selected two expression strains to move to the 1-L scale, where multiple conditions were tested (fermentation confirmation). The fermentations were sampled at multiple time points and analyzed using BLI (Figure 3). From those fermentation activities, optimal fermentation conditions that were suitable for scaling to larger volumes were identified.
Rapid Protein Analysis
In its extensive history of protein expression and product development, Pfenex has screened hundreds of thousands of expression strains and successfully delivered high-quality, active and soluble product for multiple different classes of proteins including monoclonal antibodies, antibody derivatives (Fab, Fab′, single-chain antibodies, etc.), growth factors, cytokines, and vaccine antigens. As part of that extensive expression strain screening history, Pfenex has developed and implemented several BLI assays, for rapid protein analysis. The Octet platform provides a method for measuring protein titer and verifying functionality in a simple and direct binding assay format that is broadly applicable across a variety of protein types.
Author Details
Greg Cantin, PhD, is group leader, analytical biochemistry, at Pfenex, Inc., 10790 Roselle Street, San Diego, CA, 92121; 1-858-352-4400; [email protected].
REFERENCES
1.) Retallack, DM. 2007. Transport of Heterologous Proteins to the Periplasmic Space of Pseudomonas fluorescens Using a Variety of Native Signal Sequences. Biotechnol. Lett. 29:1483-1491.
2.) Squires, CH 2011. A Toolbox for Protein Expression Scientists. Innov. Pharma. Technol.:68-72.
3.) Retallack, DM, H Jin, and L Chew. 2012. Reliable Protein Production in a Pseudomonas fluorescens Expression System. Protein Expr. Purif. 81:157-165.
4.) Concepcion, J. 2009. Label-Free Detection of Biomolecular Interactions Using BioLayer Interferometry for Kinetic Characterization. Comb. Chem. High Throughput Screen 12:791-800.
5.) Jin, H. 2011. Soluble Periplasmic Production of Human Granulocyte Colony- Stimulating Factor (G-CSF) in Pseudomonas fluorescens. Protein Expr. Purif. 78:69-77.
6.) Herman, AC, TC Boone, and HS Lu. 1996. Characterization, Formulation, and Stability of Neupogen (Filgrastim): A Recombinant Human Granulocyte-Colony Stimulating Factor. Pharm. Biotechnol. 9:303-328.