Voices of Biotech
Podcast: MilliporeSigma says education vital to creating unbreakable chain for sustainability
MilliporeSigma discusses the importance of people, education, and the benefits of embracing discomfort to bolster sustainability efforts.
June 1, 2012
Microorganisms play a vital role in modern life — with applications ranging from wine fermentation to biofuel production to solutions for complex mathematical problems (1). During the past decade, microbial fermentation for protein production reached a higher level of sophistication and wider adoption. When BPI was first published in 2003, the physical and biological characteristics of many microbial cells and the attributes of their fermentation processes were well known. Nonetheless, the economic environment at that time created immense pressure on the biopharmaceutical industry to drive innovation and emphasize manufacturing efficiency (2).
BPI’s Protein Expressions supplement in 2004 reviewed microbial fermentation, its advantages over mammalian cell expression (e.g., lower generation time, growth time, media costs, robustness), and its shortfalls (e.g., for most systems, glycosylation and posttranslational modifications) (3). Our 2008 coverage of microbial expressions confirmed that companies continued to use those same systems, typically prokaryotes such as bacteria (e.g., Escherichia coli), eukaryotes such as filamentous fungi (e.g., Aspergillus spp.) and yeast (e.g. Saccharomyces cerevisiae and Pichia pastoris) (4). At that time, of the recombinant proteins produced in the United States and Europe, 55% were expressed using microbes — 40% bacteria (39% of which as a form of Escherichia coli) and 15% yeast (5). Also at that time, however, a few other promising microbial systems had begun to gain attention along with studies involving novel applications, “platform” technologies, and improved analytical and monitoring techniques.
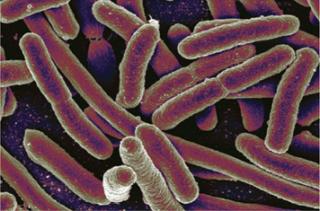
Escherichia coli ()
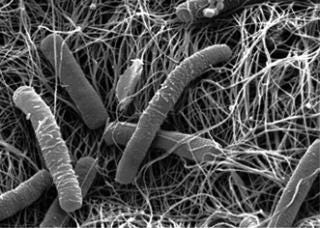
Escherichia coli ()
Escherichia coli
The most prominent expression system after Chinese hamster ovary (CHO) cells is E. coli. Especially robust and economical for the production of antibodies and recombinant proteins, it continues to be the top microbial host (of more than 30 available options) (6). Studies have demonstrated the ability of E. coli to make milligram-quantities of glycosylated proteins, thereby broadening its potential applications (7).
During the past decade, new E. coliplatform technologies have been developed. For example, in 2009, Mücke et al. demonstrated a successful technology based on E. coli secretion and high-cell-density fermentation for producing high yields of human antibody fragments (Fabs) (8). Results showed that yields were 40× higher using secretion technology than were those from periplasmic microbial expression systems. A separate study on Fabs expressed using Pichia pastoris resulted in mixed results when compared with expression from a CHO platform (9). Also in 2009, Pattnaik et al. described the application of membrane technology in the production of inclusion-body proteins from E. coli (10). They used a “highly productive fermentation process and high-throughput purification” to successfully produce (and scale) therapeutic proteins from E. coli.
Industrial applications of E. coli host systems began in the 1980s and continue to be core business strengths for many companies. Boehringer Ingelheim Austria GmbH, for example, claims to have “pioneered the microbial fermentation and purification of therapeutic proteins from E. coli since 1982.” (11). Sandoz was also an early (1980) developer of the recombinant protein interferon alpha at an industrial-scale fermentation. And Life Technologies has developed its BL21 Star E. coli expression strain for use with its pET expression system.
In 2008, more than 70% of Lonza’s fermentation projects were on an E. coli platform (with P. pastoris gaining preference) (12). Since then, the company has developed its XS Microbial Expression Technologies platform and expanded its E. coli and yeast expression technologies into developing and manufacturing plasmid DNA, antibody fragments, and protein scaffolds. Other major companies providing microbial fermentation capabilities include SAFC and Biocon. Yeast
Yeast expression platforms are attractive because of their well-defined genetic properties, ease of genetic manipulation, and simple fermentation processes that promote rapid growth rates. In bioprocessing, methylotropic yeast platforms are best known for their commercial production of recombinant proteins (13).Fermentation times are typically longer than those for E. coli but shorter than for mammalian cell cultures. The organisms have been shown not to harbor pathogens, viral inclusions, or pyrogens, so downstream processing is simpler than that of mammalian cell culture. As eukaryotes, fungi can carry out posttranslational folding such that engineered strains can execute complex N-glycosylation structures. However, their pattern of glycosylation has been reported to differ significantly from that of natural human proteins (14).
Saccharomyces cerevisiae, or “bakers yeast,” was the first yeast expression system. S. cerevisiae does not secrete many homologous proteins during growth, so its growth medium is mainly composed of heterologous proteins. Its biopharmaceutical uses include production of insulin, hepatitis B vaccines, and antibody fragments. Future applications of this yeast may include helping researchers study the effects of specific genomic changes and a gene’s function, ultimately leading toward more personalized medicine (15). During the past decade, several resources for S. cerevisiae research have grown, most notably the Saccharomyces Genome Database (SGD, www.yeastgenome.org).
Pichia pastoris is a unicellular methylotropic yeast capable of providing prokaryotic growth characteristics and eukaryotic-like posttranslational protein modifications. P. pastoris expression continues to gain popularity in the production of vaccines, antibody fragments, hormones, cytokines, matrix proteins, and biosimilars. During the past 10 years, nearly 1,800 articles have been published that center on the P. pastoris expression system (PubMed). The system has shown advantages over E. coli expression such as for human adiponectin (a promising therapy for metabolic syndromes) (16). The platform also has been shown to pr
oduce binding affinities and in vitro inhibition properties comparable to those of CHO cell systems. Kunert et al. compared the expression of an antibody fragment using CHO and P. pastoris systems. Their results showed that compared with CHO, development time is faster and less expensive (8). Emerging Microbial Host Systems
The past decade saw the introduction and growing interest of other expression technologies. Here are two we shall keep an eye on during our next 10 years.
Pseudomonas fluorescens: In 2009, Dow announced formation of a new company (Pfenex) based on its Pfēnex Expression Technology system. The Pseudomonas fluorescens-based platform uses “high-throughput, parallel processing methodologies for optimized protein production” (17). Since then, the product pipeline already contains five candidates, including biosimilars, biodefense molecules, and vaccine antigens. The company published cost comparisons with E. coli and P. pastoris expression systems using “a computer-aided approach to process economics” to show costs associated with the production of an aglycosylated protein (18).
Lactoccocus latis: This system has shown potential as “a viable choice for membrane proteins” (9). Traditionally used in food fermentation, L. latis is a gram-positive lactic bacterium that is “now used widely for large-scale overproduction of heterologously expressed proteins. Recombinant membrane proteins can be produced with affinity tags for efficient detection and purification from crude membrane protein extracts” (19). Fermentation Monitoring Systems
The fermentation process must be closely monitored to assess the health of the solution and measure cell concentrations for determining time for optimum yield. Methods based on near-infrared (NIR) spectroscopy sensors and probes have been well established for measuring biomass, glycerol, glucose, ammonium, acetate, and lactate (20). Current efforts continue in establishing at-line and on-line technologies and the use of design of experiments (DoE) strategies to assess the robustness of the fermentation process (21)
Advances in monitoring systems and enumerating technologies have also taken place this past decade. BD, for example, developed its FACSMicroCount system, and scientists from Hamburg University of Applied Sciences have a biomass monitoring system based on radiofrequency impedance (22).
Other advances in fermentation include development of animal-free media components (e.g., peptones) and the incorporation of flexible solutions in fermentation, including the use of disposable components. Single-use bioreactors for both mammalian cells and microbial fermentation have been developed that integrate sensors and provide feedback through various display units (e.g., Cell-tainer bioreactor from CELL-ution Biotech BV).
About the Author
Author Details
Maribel Rios is managing editor of BioProcess International; [email protected].
1.) Johnston, R. 2010. The Dinosaurs Reborn: Evaluating Stainless Steel and Disposables in Large-Scale Biomanufacturing. BioProcess Int. 8:28-33.
2.) Chatrathi, K. 2004. Metabolic Cooking Capacity of Fermentors. BioProcess Int. 2:44-48.
3.) Pelin, K, K Phillips, and V. Sarantschin. 2003. Building a GMP Bacterial and Fungal Fermentation Facility. BioProcess Int. 1:56-60.
4.) Mirro, R, and K. Voll. 2009. Which Impeller is Right for Your Cell Line?. BioProcess Int. 7:52-57.
5.) Dhanasekharan, K. 2006. Design and Scale-Up of Bioreactors Using Computer Simulation. BioProcess Int. 4:34-42.
6.) Julien, C, and W. Whitford. 2007. Getting the Most from Your Bioreactor. BioProcess Int. 5:S4-S10.
7.) Julien, C, and W. Whitford. 2007. Bioreactor Monitoring, Modeling, and Simulation. BioProcess Int. 5:S10-S17.
8.) Bonham-Carter, Jerry, and J. Shevitz. 2011. A Brief History of Perfusion Biomanufacturing. BioProcess Int. 9:24-31.
9.) Whitford, W, and JJS. Cadwell. 2009. Interest in Hollow-Fiber Perfusion Bioreactors Is Growing. BioProcess Int. 7:54-64.
10.) Acuna, J. 2011. Modeling Perfusion Processes in Biopharmaceutical Production. BioProcess Int. 9:52-59.
11.) Langer, ES. 2011. Trends in Perfusion Bioreactors. BioProcess Int. 9:18-22.
12.) Aranha, H. 2004. Disposable Systems. BioProcess Int. 2:S6-S16.
13.) Rader, R, and ES. Langer. 2012. Upstream Single-Use Bioprocessing Systems. BioProcess Int. 10:12-19.
14.) Bader, R, A Donofrio, and M. Ebling. 2005. A Disposable Mixing System for Hydrating Powdered Media and Reagents. BioProcess Int. 3:76-80.
15.) Waele, KD. 2007. A Novel Single-Use Mixing System for Buffer Preparation. BioProcess Int. 5:S56-S61.
16.) Strahlendorf, K, and K. Harper. 2010. Mixing in Small-Scale Single-Use Systems. BioProcess Int. 8:42-49.
17.) Raval, K, C-M Liu, and J. Buchs. 2006. Large-Scale Disposable Shaking Bioreactors: A Promising Choice. BioProcess Int. 4:46-50.
18.) Isailovic, B, and B. Rawlings. 2011. An Approach to Design and Performance Testing of an Impeller-Driven Single-Use Mixer. BioProcess Int. 10:60-69.
19.) Houtzager, E. 2005. Linear Scale-Up of Cell Cultures: The Next Level in Disposable Bioreactor Design. BioProcess Int. 3:60-66.
20.) Scott, C. 2007. Single-Use Bioreactors: A Brief Review of Current Technology. BioProcess Int. 5:S44-S51.
21.) Fisher, M. 2006. A Stirred-Tank Bioreactor: Delivered in Eight Weeks and One Hour. BioProcess Int. 4:S28-S30.
22.) Wilde, DD. 2009. Bridging the Gap from Reusable to Single-Use Manufacturing with Stirred, Single-Use Bioreactors. BioProcess Int. 7:S36-S41.
23.) Eibl, R, and D. Eibl. 2009. Disposable Bioreactors in Cell Culture-Based Upstream Processing. BioProcess Int. 7:S24-S27.
24.) Jia, Q. 2008. A Bioreactor System Based on a Novel Oxygen Transfer Method. BioProcess Int. 6:66-71.
25.) Rodriguez, R, J Castillo, and S. Giraud. 2010. Demonstrated Performance of a Disposable Bioreactor with an Anchorage Dependent Cell Line. BioProcess Int. 8:74-78.
26.) Li, L. 2009. A Single-Use, Scalable Perfusion Bioreactor System 7:46-54.
27.) Seamans, TC. 2008. Cell Cultivation Process Transfer and Scale-Up, Part 1. BioProcess Int. 6:26-36.
28.) Seamans, TC. 2008. Cell Cultivation Process Transfer and Scale-Up, Part 2. BioProcess Int. 6:34-42.
29.) Kinney, SD, CW Phillips, and KJ. Lin. 2007. Thermoplastic Tubing Welders and Sealers. BioProcess Int. 5:S52-S61.
30.) Boehm, J, and B. Bushnell. 2007. Providing Sterility Assurance Between Stainless Steel and Single-Use Systems. BioProcess Int. 5:S66-S71.
31.) Kane, J. 2012. Measuring kLa for Better Bioreactor Performance. BioProcess Int. 10:46-49.
32.) Mattes, RA. 2007. In Situ Monitoring of CHO Cell Culture Medium Using Near Infrared Spectroscopy. BioProcess Int. 5:S46-S50.
33.) Card, C. 2008. Near-Infrared Spectroscopy for Rapid, Simultaneous Monitoring. BioProcess Int. 6:58-66.
34.) Mattes, R. 2009. Real-Time Bioreactor Monitoring of Osmolality and pH Using Near-Infrared Spectroscopy. BioProcess Int. 7:44-50.
35.) Logan, D, and J. Carvell. 2011. A Biomass Monitor for Disposable Bioreactors. BioProcess Int. 10:48-54.
36.) Kaiser, C, JP Carvell, and RA. Luttmann. 2007. Sensitive, Compact, In Situ Biomass Measurement System: Controlling and Monitoring Microbial Fermentations Using Radio-Frequency Impedence. BioProcess Int. 5:S52-S56.
37.) Clark, K, and J. Furey. 2006. Suitability of Selected Single-Use Process Monitoring and Control Technology. BioProcess Int. 6:16-20.
38.) Furey, J, K Clark, and C. Card. 2011. Adoption of Single-Use Sensors for BioProcess Operations. BioProcess Int. 9:S36-S42.
39.) Tsai, W-L. 2012. Noninvasive Optical Sensor Technology in Shake Flasks. BioProcess Int. 10:50-56.
40.) Falkowitz, K, J Staggert, and V. Wedege. 2006. A System Approach to Improving Yields in a Disposable Bioreactor. BioProcess Int. 4:56-62.
41.) Jones, S, SD McKee, and HL Levine. 2012. Emerging Challenges in Cell Therapy Manufacturing. BioProcess Int. 10:S4-S7.
42.) Hampson, B, J Rowley, and N. Venturi. 2008. Manufacturing Patient-Specific Cell Therapy Products. BioProcess Int. 6:60-72.
43.) Rowley, J. 2012. Meeting Lot-Size Challenges of Manufacturing Adherent Cells for Therapy. BioProcess Int. 10:S16-S22
You May Also Like